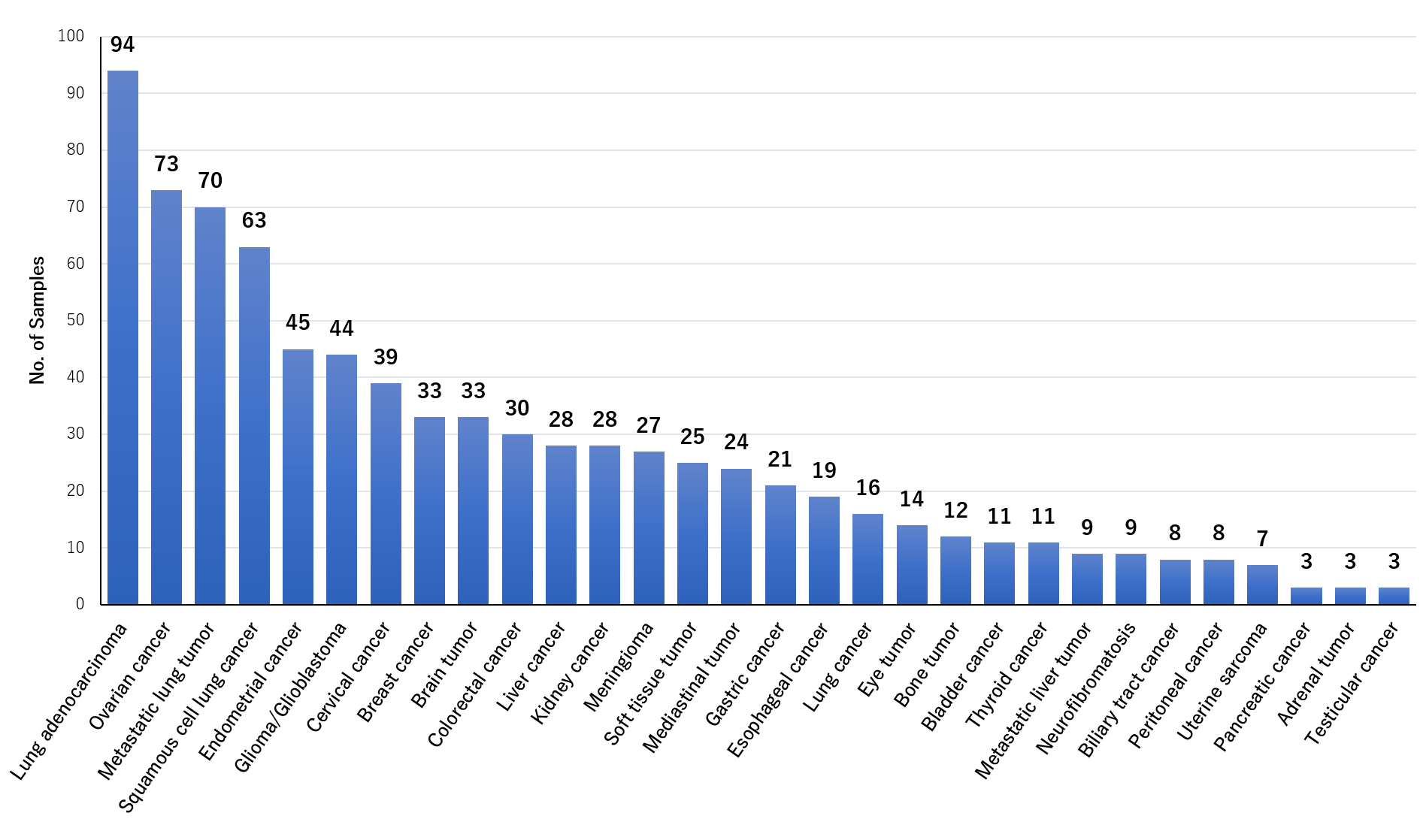Overview and Feature of Data to be provided
● Whole Exome Sequence Analysis of Japanese patients with cancer performed in Fukushima Translational Research Project.
● Tumor-Normal pair analysis providing only Tumor-Specific Variants and no personally genomic information
● Unpublished variant data
● Obtained informed consent (IC) to be able to utilize for both domestic and foreign companies
● No restriction to the purpose to utilize data
● Exclusive contract for all data to use also available
As a part of Fukushima Translational Research Project, we are performing genome analysis using tumor samples donated from Japanese cancer patients, mainly tumor-normal pair analysis with whole exome sequence 1). We have completed the analysis of 727 cases and 810 cancer samples classified into 30 cancer types(Fig. 1).
On this occasion, we announce a start to provide these data for a fee. These data have not been published yet 2). Also, informed consent (IC) has been obtained from all patients to use these data for commercial purpose. Obtained data will be utilized in companies all over the world, such as biotechnology/IT-related companies. Companies which purchased these data will be capable to use freely for any purpose. Exclusive contract for utilization of all data is also available.
Variant data will be provided in either VCF or text format. Data to be provided are obtained from tumor-normal pair analysis, therefore these are tumor-specific and do not contain personal genomic information. In addition, we also have performed Cancer Hotspot Panel analysis to all samples to make it firm detection of cancer-driver gene mutations. This information will be also provided with whole exome analysis data.
We accept the request of providing clinical information on sample corresponding to data with consultation 3). Also, we have performed transcriptome analysis in addition to whole exome analysis and these data also can be provided for a fee. If you have any questions, please contact us at the following address: tlo@ftrf.jp, or visit website: https://www.fmu.ac.jp/home/trc/about/organization/gene-factory/
1)We used Ion Ampliseq™ Exome System for analysis. Please contact us for more information.
2)The number of mutations per sample (TMB: Tumor Mutation Burden) in some tumor sample cases has been published (Cancer Immunol Immunother 2020 Jan;69(1):127-134). It has been published as numerical data, not more detailed mutation data itself.
3)Please note that we may not always be able to meet your request due to the handling of sensitive personal information.
Fig. 1 Cancer samples analyzed in this project